User Interface
Overview
When you first start PyJibe, you will be greeted with an empty user interface. To open experimental data, choose one of the corresponding options in the File menu.
Force-Distance Analysis
Force-distance (FD) measurements consist of an approach and a retract curve. The point of maximum indentation is usually defined by the acquisition software as a setpoint using an active feedback loop.
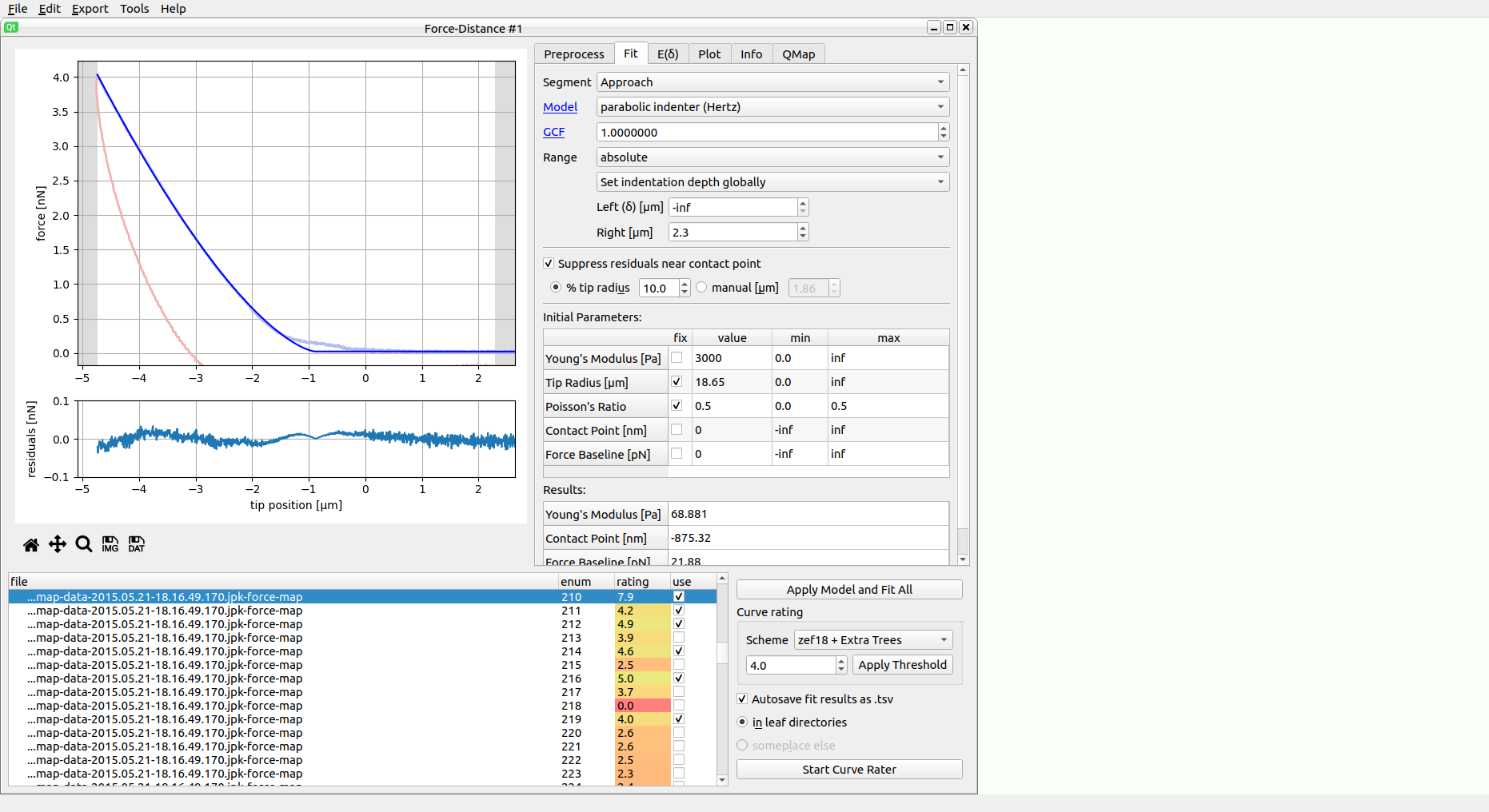
The screenshot shows a typical data analysis in PyJibe. A .jpk-force-map file has been loaded. The plot at the top shows the experimental FD curve (approach light blue, retract light red), the model fit (dark blue), and the fitting region (white region, bounded by gray background in the upper plot). Below are the fit residuals. On the right are the main controls (discussed in the sections below). At the bottom is the curve list, highlighting which curve is currently shown at the top. At the bottom right are the curve list controls.
Note
Notice how the fit residuals become small around the contact point? PyJibe comes with the feature Suppress residuals near contact point (in the fit tab on the right). This option suppresses the contributions of the FD curve near the contact point. This feature was designed for biological samples, where the physical tip-sample interactions near the “contact point” are not well-understood [MAM+19]. The contributions near the contact point are suppressed to give the remainder of the indentation curve more weight.
The dataset shown resembles a 2D scan of AFM force-distance curves from a live zebrafish spinal cord microtome section and is available on figshare [MMG20] (same data as in [MKH+20] fig. 1e and [MAM+19] fig. 3a,d,g).
Tab: Preprocess
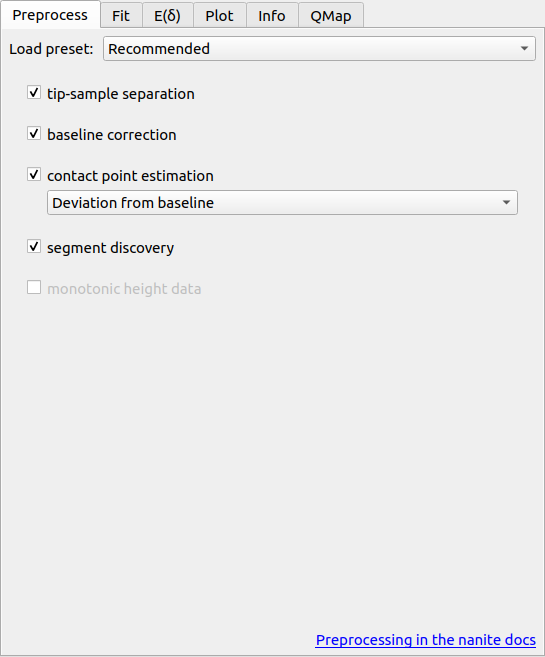
Before starting to fit FD data, it is important to perform the correct preprocessing steps. Usually, this includes computing the tip position and correcting for a force and tip position offset.
The preprocessing pipeline is defined via the checkable options in this tab. If you hover over the options, a tool tip is displayed with an explanation. If your analysis pipeline strongly depends on a good estimate of the contact point, then you have multiple options which are described in detail in the nanite docs.
Tab: Fit
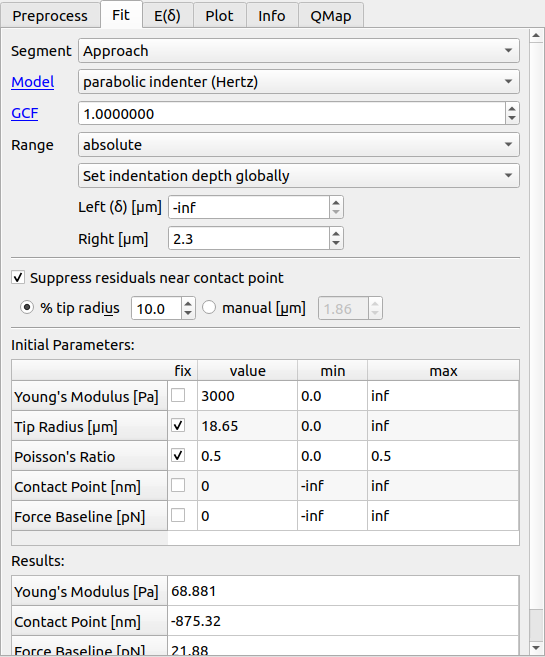
PyJibe offers a variety of options for fitting FD data. In the upper part, you may select the segment (approach or retract), the axes, the fit model, and the fitting range.
- Range:
The fitting range can be set to absolute (relative to the X axis after preprocessing) or relative to the contact point. In the latter case, the fit is repeated four times, updating the new contact point at each iteration. You have three option for setting the indentation depth (left part of the fit interval):
globally: For each FD curve, the same indentation depth is used (after preprocessing has been applied).
individually: You select the indentation depth for each FD curve separately.
guess optimal indentation depth: The optimal indentation depth is guessed using the E(δ)-curve.
- Suppress residuals near contact point:
This option multiplies the residuals with a linear ramp, left and right of the contact point. The ramp has a value of 0 at the contact point and a value of 1 at the distance defined by the controls. All other residuals or not changed. This feature was designed for biological samples, where the physical tip-sample interactions near the “contact point” are not well-understood [MAM+19]. The contributions near the contact point are suppressed to give the remainder of the indentation curve more weight.
- Ancillary Parameters:
Ancillary parameters (not shown in the screenshot here), are parameters that are defined in the fitting model. They are computed prior to the fit and can be set as fitting parameters. Standard models usually do not have ancillary parameters.
- Initial Parameters:
These are the parameters set initially during fitting. In the table, you may choose which parameters should remain fixed during fitting, what the initial values should be, and in which interval this value may be varied.
- Results:
The fit results of the varied parameters are shown here.
Tab: E(δ)
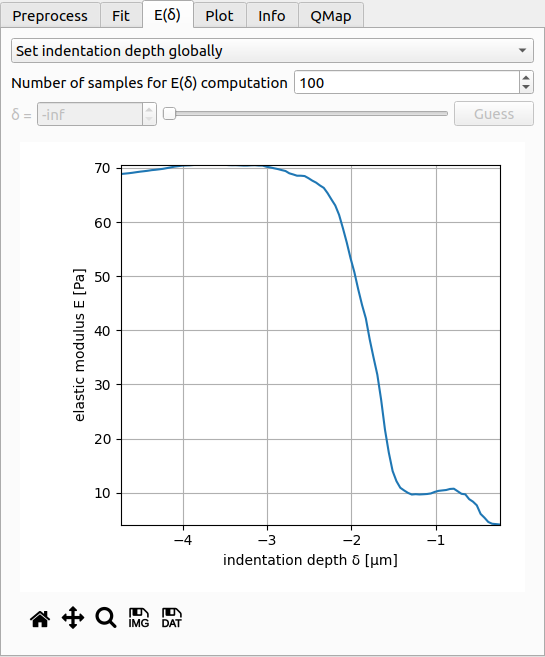
The E(δ) curve is used to test whether the fitted Young’s modulus E is dependent on the fitting interval. For a reliable fit, the E(δ) curve should exhibit a plateau. This is the case for the present example, where the fitted value of E does not vary much in the range from δ=-2.5µm to δ=-4.8µm (maximum indentation).
The control at the top is identical to the control in the fitting tab. You may choose the number of samples of the E(δ). The controls for setting the indentation depth manually are enabled when you select set indentation depth individually in the dropdown menu. Below the plot you have the options to export the E(δ) curve as an image or as a data file (.tsv). If you would like to export all E(δ) curves of the dataset, you can do so via the Export menu of the main window.
Tab: Plot
The plotting tab allows you to modify the plot settings. By default, the plot is adapted to the fitting interval. You may, however, modify the axes ranges manually by unchecking the check boxes.
Tab: Info
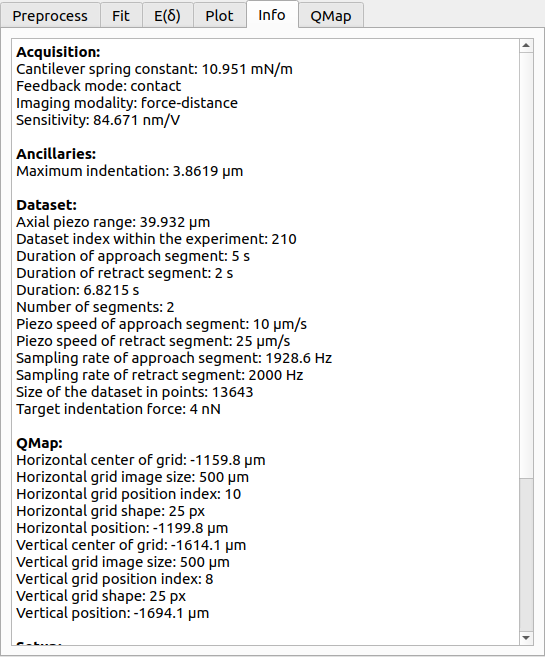
The info tab shows metadata related to the currently shown curve. Unknown values are indicated as nan.
- Acquisition:
Settings of the acquisition software
- Ancillaries:
Ancillary parameters are computed for each model. Here, the maximum indentation is listed, which is the difference between the fitted contact point and the value of the tip position where the fitted curve has its maximum. Fit models may have their own specific ancillary parameters.
- Dataset:
Measurement parameters of the dataset
- QMap:
If the curve is part of a quantiative map (2D FD scan), then the scan grid properties and the curve position are listed.
- Setup:
Information about the AFM setup used
- Storage:
Information about the data file, which are not particularly important for the experiment, but allow to identify a dataset or curve
Tab: QMap
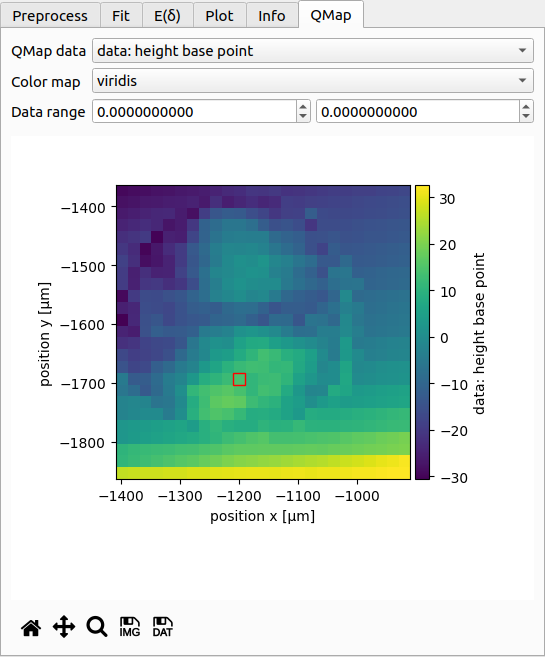
If the dataset consists of a quantitative map, then this map is shown here. It serves as an interactive overview of the dataset. You can choose which data to plot (e.g. piezo height, fitted parameters, curve rating, scan order), which colormap to use, and how the colormap should be scaled. The coordinate axes are identical to those shown in the info tab.
The red square in the plot indicates which curve is currently shown. You may click on the plot to select curves manually (as opposed to using the curve list at the bottom of the window).
Below the plot are controls for exporting the qmap as an image or as a text file.
Curve list controls
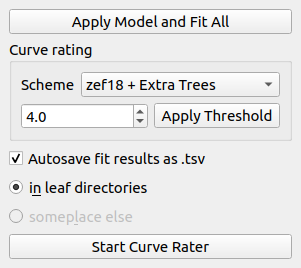
The curve list controls perform operations on the entire FD curve list. The Apply Model and Fit All button does exactly what it says.
Curve rating allows you to rate each indentation curve. This is useful e.g. when you would like to sort out bad curves automatically. In this example, all curves with a rating below a threshold of 4.5 are excluded from further analysis using the zef18 + Extra Trees rating scheme [MAM+19].
Below the curve rating options, you may enable or disable autosaving of the fit results. Only the results for curves that are selected as “use” in the curve list are exported.
For more information on rating, please have a look at the nanite documentation. The button at the bottom starts the PyJibe curve rater which is compatible to the nanite rating workflow. If you would like to import your own training set, please read the quick guide Importing a pre-computed training set.
Advanced options
Extensions
Under Edit | Preferences | Extensions, you may import your own custom fit models. See Loading Extensions (nanite fit models) for more information.
Expert and Developer mode
Under Edit | Preferences | Advanced, you can enable expert and developer mode.
Expert mode:
Enables you to manually set the minimizer method and the corresponding keyword arguments applied by lmfit.
Developer mode:
Everything that is available in expert mode.
Allows you to open experimental data that were not recorded via force-distance modality (e.g. creep-compliance). Related issues are nanite #11 and afmformats #15.
Slows down loading of large datasets (because the modality has to be determined first).
Adds the force-distance fitting model sneddon_spher to the list of available fit models (it is excluded by default, because it is virtually identical to the sneddon_spher_approx model which is much faster).
Displays and exports hidden model parameters (parameters whose name starts with an underscore
_).